Prerequisites
It is recommended that you familiarise yourself with R first by sitting our Introduction to R tutorial.
It also requires that you have the DataSHIELD training environment installed on your machine, see our Installation Instructions for Linux, Windows, or Mac.
Help
DataSHIELD support is freely available in the DataSHIELD forum by the DataSHIELD community. Please use this as the first port of call for any problems you may be having, it is monitored closely for new threads.
DataSHIELD bespoke user support and also user training classes are offered on a fee-paying basis. Please enquire at datashield@newcastle.ac.uk for current prices.
Introduction
This is the third in a 6-part DataSHIELD tutorial series. Please see below for links to all prior and subsequent parts:
Introduction and logging in
Basic statistics and data manipulations
Assign functions and tables
Plotting graphs
Subsetting
Modelling
Quick reminder for logging in:
- Follow instructions to Start the Opal VMs.
Recall from the installation instructions, the Opal web interface is a simple check to tell if the VMs have started. Load the following urls, waiting at least 1 minute after starting the training VMs.
Start R/RStudio
Load Packages
#load libraries library(DSI) library(DSOpal) library(dsBaseClient)
Build your login dataframe
builder <- DSI::newDSLoginBuilder()
builder$append(server = "study1", url = "http://192.168.56.100:8080/",
user = "administrator", password = "datashield_test&",
table = "CNSIM.CNSIM1", driver = "OpalDriver")
builder$append(server = "study2", url = "http://192.168.56.101:8080/",
user = "administrator", password = "datashield_test&",
table = "CNSIM.CNSIM2", driver = "OpalDriver")
logindata <- builder$build()
connections <- DSI::datashield.login(logins = logindata, assign = TRUE, symbol = "D")
- Command to logout:
DSI::datashield.logout(connections)
Sub-setting
Limitations on subsetting
Sub-setting is particularly useful in statistical analyses to break down variables or tables of variables into groups for analysis. Repeated sub-setting, however, can lead to thinning of the data to individual-level records that are disclosive (e.g. the statistical mean of a single value point is the value itself). Therefore, DataSHIELD does not subset an object below the minimal subset length set by the data providers (typically this is ≤ 4 observations).
In DataSHIELD there are currently 3 functions that allow us to generate subset data:
- ds.subsetByClass
- ds.dataFrameSubset
- ds.subset (WARNING: this function will be deprecated in the release of 6.1, all functionallity has been added to ds.dataFrameSubset which will become the one-stop replacement).
Sub-setting using ds.subsetByClass
- The
ds.subsetByClassfunction generates subsets for each level of acategoricalvariable. If the input is a data frame it produces a subset of that data frame for each class of each categorical variable held in the data frame. - Best practice is to state the categorical variable(s) to subset using the
variablesargument, and the name of the subset data using thesubsetsargument. - The example subsets
GENDERfrom our assigned data frameD, the subset data is namedGenderTables:
ds.subsetByClass(x = 'D', subsets = "GenderTables", variables = 'GENDER', datasources = connections)
- The output of
ds.subsetByClassis held in alistobject stored server-side, as the subset data contain individual-level records. If no name is specified in thesubsetsargument, the default name "subClasses"is used.
Running ds.subsetByClass on a data frame without specifying the categorical variable in the argument
variables
will create a subset of all categorical variables. If the data frame holds many categorical variables the number of subsets produces might be too large - many of which may not be of interest for the analysis.
In the previous example, the
GENDER
variable in assigned data frame
D
had females coded as 0 and males coded as 1. When
GENDER
was subset using the ds.subsetByClass function, two subset tables were generated for each study dataset; one for females and one for males.
- The
ds.namesfunction obtains the names of these subset data:
ds.names('GenderTables', datasources = connections)
Aggregated (exists("GenderTables")) [==================================================] 100% / 0s
Aggregated (classDS("GenderTables")) [=================================================] 100% / 0s
Aggregated (namesDS(GenderTables)) [===================================================] 100% / 0s
$study1
[1] "GENDER.level_0" "GENDER.level_1"
$study2
[1] "GENDER.level_0" "GENDER.level_1"
Sub-setting using ds.subset
This function is soon to be deprecated. Its replacement will be ds.dataFrameSubset().
ds.dataFrameSubset() uses very different arguments to ds.subset()
Changes will be coming soon to this page. Use function help to investigate how ds.dataFrameSubset() works similarly.
The function
ds.subset
allows general sub-setting of different data types e.g. categorical, numeric, character, data frame, matrix. It is also possible to subset rows (the individual records). No output is returned to the client screen, the generated subsets are stored in the server-side R session.
- The example below uses the function
ds.subsetto subset the assigned data frameDby rows (individual records) that have no missing values (missing values are denoted withNA) given by the argumentcompleteCases=TRUE. The output subset is named "D_without_NA":
ds.subset(x='D', subset='D_without_NA', completeCases=TRUE, datasources = connections)
The ds.subset function prints an invalid message to the client screen to inform if missing values exist in a subset.
#In order to indicate that a generated subset dataframe or vector is invalid all values within it are set to NA!
An invalid message also denotes subsets that contain less than the minimum cell count determined by data providers.
- The second example creates a subset of the assigned data frame
Dwith BMI values ≥ 25 using the argumentlogicalOperator. The subset object is namedBMI25plususing thesubsetargument and is not printed to client screen but is stored in the server-side R session:
ds.subset(x='D', subset='BMI25plus', logicalOperator='PM_BMI_CONTINUOUS>=', threshold=25, datasources = opals)
The subset of data retains the same variables names i.e. column names
- To verify the subset above is correct (holds only observations with BMI ≥ 25) the function
ds.quantileMeanwith the argumenttype='split'will confirm the BMI results for each study are ≥ 25.
ds.quantileMean('BMI25plus$PM_BMI_CONTINUOUS', type='split', datasources = opals)
$`dstesting-100`
5% 10% 25% 50% 75% 90% 95% Mean
25.3500 25.7100 27.1500 29.2000 32.0600 34.6560 36.4980 29.9019
$`dstesting-101`
5% 10% 25% 50% 75% 90% 95% Mean
25.46900 25.91800 27.19000 29.27000 32.20500 34.76200 36.24300 29.92606
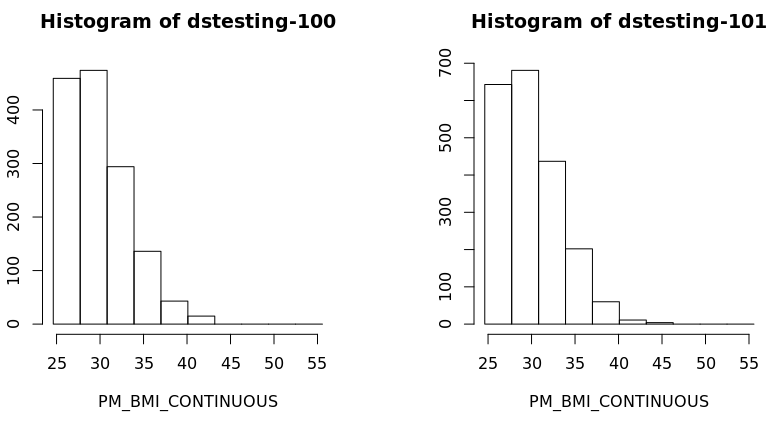
- Also a histogram of the variable BMI of the new subset data frame could be created for each study separately:
ds.histogram('BMI25plus$PM_BMI_CONTINUOUS', datasources = opals)
Warning: dstesting-100: 2 invalid cells
Warning: dstesting-101: 1 invalid cells
[[1]]
$breaks
[1] 23.93659 27.17016 30.40373 33.63731 36.87088 40.10445 43.33803 46.57160 49.80518 53.03875 56.27232
$counts
[1] 365 511 331 150 49 15 0 0 0 0
$density
[1] 0.079212771 0.110897880 0.071834047 0.032553194 0.010634043 0.003255319 0.000000000 0.000000000 0.000000000 0.000000000
$mids
[1] 25.55337 28.78695 32.02052 35.25409 38.48767 41.72124 44.95482 48.18839 51.42196 54.65554
$xname
[1] "xvect"
$equidist
[1] TRUE
attr(,"class")
[1] "histogram"
[[2]]
$breaks
[1] 23.93659 27.17016 30.40373 33.63731 36.87088 40.10445 43.33803 46.57160 49.80518 53.03875 56.27232
$counts
[1] 506 750 476 229 62 11 4 0 0 0
$density
[1] 0.0767450721 0.1137525773 0.0721949690 0.0347324536 0.0094035464 0.0016683711 0.0006066804 0.0000000000 0.0000000000 0.0000000000
$mids
[1] 25.55337 28.78695 32.02052 35.25409 38.48767 41.72124 44.95482 48.18839 51.42196 54.65554
$xname
[1] "xvect"
$equidist
[1] TRUE
attr(,"class")
[1] "histogram
Conclusion
The next part in this DataSHIELD tutorial series is:
Introduction and logging in
Basic statistics and data manipulations
Assign functions and tables
Plotting graphs
Subsetting
Modelling
Also remember you can:
- get a function list for any DataSHIELD package and
- view the manual help page individual functions
- in the DataSHIELD test environment it is possible to print analyses to file (.csv, .txt, .pdf, .png)
- take a look at our FAQ page for solutions to common problems such as Changing variable class to use in a specific DataSHIELD function.
- Get support from our DataSHIELD forum.