It is recommended that you familiarise yourself with R first by sitting our Session 1: Introduction to R tutorial |
Datashield user support is provided via the DataSHIELD Forum, DataSHIELD user training classes are offered on a fee basis. Please enquire at datashield@newcastle.ac.uk for current prices. |
This tutorial introduces users R/RStudio users to DataSHIELD commands and syntax. Throughout this document we refer to R, but all commands are run in the same way in Rstudio. This tutorial contains a limited number of examples; further examples are available in each DataSHIELD function manual page that can be accessed via the help function.
DataSHIELD commands call functions that range from carrying out pre-requisite tasks such as login to the data providers, to generating basic descriptive statistics, plots and tabulations. More advance functions allow for users to fit generalized linear models and generalized estimating equations models. R can list all functions available in DataSHIELD.
This section explains the functions we will call during this tutorial. Although this knowledge is not required to run DataSHIELD analyses it helps to understand the output of the commands. It can explain why some commands call functions that return nothing to the user, but rather store the output on the server of the data provider for use in a second function.
In DataSHIELD the person running an analysis (the client) uses
client-side functions
to issue commands (instructions). These commands initiate the execution (running) of
server-side functions
that run the analysis server-side (behind the firewall of the data provider). There are two types of server-side function:
assign functions
and
aggregate functions
.
|
Follow the instructions depending on your training environment:
Your trainer will have started your Opal training servers in the cloud for you.
Your trainer will give you the IP address of the DataSHIELD client portal ending :8787
They will also provide you with a username and password to login with.
The DataSHIELD cloud training environment does not use fixed IP addresses, the client and opal training server addresses change each training session. As part of the user tutorial you learn how to build a DataSHIELD login dataframe. In a real world instance of DataSHIELD this is populated with secure certificates not text based usernames and passwords. |
Ctrl+Enterserver <- c("ds-training-server1", "ds-training-server2") #The server names
url <- c("http://XXX.XXX.XXX.XXX:8080", "http://XXX.XXX.XXX.XXX:8080") # These IP addresses change
user <- "administrator"
password <- "datashield_test&"
table <- c("CNSIM.CNSIM1","CNSIM.CNSIM2") #The data tables used in the tutorial
my_logindata <- data.frame(server,url,user,password,table) |
my_logindata |
Start your Opal Training VMs.
As part of the user tutorial you learn how to build a DataSHIELD login dataframe. In a real world instance of DataSHIELD this is populated with secure certificates not text based usernames and passwords. |
server <- c("dstesting-100", "dstesting-101") #The VM names
url <- c("http://192.168.56.100:8080", "http://192.168.56.101:8080") # The fixed IP addresses of the training VMs
user <- "administrator"
password <- "datashield_test&"
table <- c("CNSIM.CNSIM1","CNSIM.CNSIM2") #The data tables used in the tutorial
my_logindata <- data.frame(server,url,user,password,table) |
my_logindata |
opal to login and logout
dsBaseClient ,
dsStatsClient ,
ds.GraphicsClient and
dsModellingClient containing all DataSHIELD functions referred to in this tutorial.
library function into the command line as given in the example below:#load libraries library(opal) library(dsBaseClient) library(dsStatsClient) library(dsGraphicsClient) library(dsModellingClient) |
The output in R/RStudio will look as follows:
library(opal) #Loading required package: RCurl #Loading required package: bitops #Loading required package: rjson library(dsBaseClient) #Loading required package: fields #Loading required package: spam #Loading required package: grid library(dsStatsClient) library(dsGraphicsClient) library(dsModellingClient) |
You might see the following status message that you can ignore. The message refers to the blocking of functions within the package. |
opals that calls the
datashield.login function to log into the desired Opal servers. In the DataSHIELD test environment
logindata is our login template for the test Opal servers.opals <- datashield.login(logins=my_logindata,assign=TRUE) |
The output below indicates that each of the two Opal training servers
study1 and
study2 contain the same 11 variables listed in capital letters under
Variables assigned: .
opals <- datashield.login(logins=my_logindata,assign=TRUE) # Logging into the collaborating servers # No variables have been specified. # All the variables in the opal table # (the whole dataset) will be assigned to R! # Assigining data: # study1... # study2... # Variables assigned: # study1--LAB_TSC, LAB_TRIG, LAB_HDL, LAB_GLUC_ADJUSTED, PM_BMI_CONTINUOUS, DIS_CVA, MEDI_LPD, DIS_DIAB, DIS_AMI, GENDER, PM_BMI_CATEGORICAL # study2--LAB_TSC, LAB_TRIG, LAB_HDL, LAB_GLUC_ADJUSTED, PM_BMI_CONTINUOUS, DIS_CVA, MEDI_LPD, DIS_DIAB, DIS_AMI, GENDER, PM_BMI_CATEGORICAL |
In Horizontal DataSHIELD pooled analysis the data are harmonized and the variables given the same names across the studies, as agreed by all data providers. |
The datashield.login function from the R package All the commands sent after login are processed within the server-side R instance only allows a specific set of commands to run (see the details of a typical horizontal DataSHIELD process). The server-side R session is wiped after loging out. If we do not specify individual variables to assign to the server-side R session, all variables held in the Opal servers are assigned. Assigned data are kept in a data frame named |
Users can specify individual variables to assign to the server-side R session. It is best practice to first create a list of the Opal variables you want to analyse.
myvar
that lists the Opal variables required for analysis:
LAB_HDL
and
GENDER
variables
argument in the function
datashield.login
uses
myvar
, which then will call only this list.myvar <- list('LAB_HDL', 'GENDER')
opals <- datashield.login(logins=my_logindata,assign=TRUE,variables=myvar)
#Logging into the collaborating servers
#Assigining data:
#study1...
#study2...
#Variables assigned:
#study1--LAB_HDL, GENDER
#study2--LAB_HDL, GENDER
|
Assigned data are kept in a data frame (table) named |
symbol
in the
datashield.login
function to change the name of the data frame from
D
to
mytable
.myvar <- list('LAB_HDL', 'GENDER')
opals <- datashield.login(logins=my_logindata,assign=TRUE,variables=myvar, symbol='mytable')
#Logging into the collaborating servers
#Assigining data:
#study1...
#study2...
#Variables assigned:
#study1--LAB_HDL, GENDER
#study2--LAB_HDL, GENDER
|
Only DataSHIELD developers will need to change the default value of the last argument, |
Almost all functions in DataSHIELD can display split results (results separated for each study) or pooled results (results for all the studies combined). This can be done using the |
It is possible to get some descriptive or exploratory statistics about the assigned variables held in the server-side R session such as number of participants at each data provider, number of participants across all data providers and number of variables. Identifying parameters of the data will facilitate your analysis.
D
can be found using the
ds.dim
command in which
type='split'
is the default argument:opals <- datashield.login(logins=my_logindata,assign=TRUE) ds.dim(x='D') |
The output of the command is shown below. It shows that in study1 there are 2163 individuals with 11 variables and in study2 there are 3088 individuals with 11 variables:
> opals <- datashield.login(logins=my_logindata,assign=TRUE) Logging into the collaborating servers No variables have been specified. All the variables in the opal table (the whole dataset) will be assigned to R! Assigning data: study1... study2... Variables assigned: study1--LAB_TSC, LAB_TRIG, LAB_HDL, LAB_GLUC_ADJUSTED, PM_BMI_CONTINUOUS, DIS_CVA, MEDI_LPD, DIS_DIAB, DIS_AMI, GENDER, PM_BMI_CATEGORICAL study2--LAB_TSC, LAB_TRIG, LAB_HDL, LAB_GLUC_ADJUSTED, PM_BMI_CONTINUOUS, DIS_CVA, MEDI_LPD, DIS_DIAB, DIS_AMI, GENDER, PM_BMI_CATEGORICAL > ds.dim(x='D') $study1 [1] 2163 11 $study2 [1] 3088 11 |
type='combine'
argument in the
ds.dim
function to identify the number of individuals (5251) and variables (11) pooled across all studies:ds.dim('D', type='combine')
#$pooled.dimension
#[1] 5251 11
|
ds.colnames
function on the assigned data frame
D
:ds.colnames(x='D') #$study1 # [1] "LAB_TSC" "LAB_TRIG" "LAB_HDL" "LAB_GLUC_ADJUSTED" "PM_BMI_CONTINUOUS" "DIS_CVA" # [7] "MEDI_LPD" "DIS_DIAB" "DIS_AMI" "GENDER" "PM_BMI_CATEGORICAL" #$study2 # [1] "LAB_TSC" "LAB_TRIG" "LAB_HDL" "LAB_GLUC_ADJUSTED" "PM_BMI_CONTINUOUS" "DIS_CVA" # [7] "MEDI_LPD" "DIS_DIAB" "DIS_AMI" "GENDER" "PM_BMI_CATEGORICAL" |
ds.class
function to identify the class (type) of a variable - for example if it is an integer, character, factor etc. This will determine what analysis you can run using this variable class. The example below defines the class of the variable LAB_HDL held in the assigned data frame D, denoted by the argument
x='D$LAB_HDL'
ds.class(x='D$LAB_HDL') #$study1 #[1] "numeric" #$study2 #[1] "numeric" |
As LAB_HDL is a numeric variable the distribution of the data can be explored.
ds.quantileMean
returns the quantiles and the statistical mean. It does not return minimum and maximum values as these values are potentially disclosive (e.g. the presence of an outlier). By default
type='combined'
in this function - the results reflect the quantiles and mean pooled for all studies. Specifying the argument
type='split'
will give the quantiles and mean for each study:ds.quantileMean(x='D$LAB_HDL') # Quantiles of the pooled data # 5% 10% 25% 50% 75% 90% 95% Mean # 1.042161 1.257601 1.570058 1.901810 2.230529 2.521782 2.687495 1.561957 |
ds.mean
use the argument
type
to request split results:ds.mean(x='D$LAB_HDL') #$`Global mean` #[1] 1.561957 |
So far all the functions in this section have returned something to the screen. Some functions (assign functions) create new objects in the server-side R session that are required for analysis but do not return an anything to the client screen. For example, in analysis the log values of a variable may be required.
ds.log
computes the natural logarithm. It is possible to compute a different logarithm by setting the argument
base
to a different value. There is no output to screen:ds.log(x='D$LAB_HDL') |
'LAB_HDL_log'
)
newobj
argument:ds.log(x='D$LAB_HDL', newobj='LAB_HDL_log') |
D
). We can check the size of the new LAB_HDL_log vector we generated above; the command should return the same figure as the number of rows in the data frame 'D'.ds.length (x='LAB_HDL_log') #$total.number.of.observations #[1] 5251 ds.length(x='D$LAB_HDL') #$total.number.of.observations #[1] 5251 |
The |
ds.assign
we subtract the pooled mean calculated earlier from LAB_HDL (mean centring) and assign the output to a new variable called
LAB_HDL.c
. The function returns no output to the client screen, the newly created variable is stored server-side.ds.assign(toAssign='D$LAB_HDL-1.562', newobj='LAB_HDL.c') |
Further DataSHIELD functions can now be run on this new mean-centred variable
LAB_HDL.c
. The example below calculates the mean of the new variable
LAB_HDL.c
which should be approximately 0.
ds.mean(x='LAB_HDL.c') #$`Global mean` #[1] -4.280051e-05 |
The function
ds.table1D
creates a one-dimensional contingency table of a categorical variable. The default is set to run on pooled data from all studies, to obtain an output of each study set the argument
type='split'
.
GENDER
. The function returns the counts and the column and row percent per category, as well as information about the validity of the variable in each study dataset:ds.table1D(x='D$GENDER') # $counts # D$GENDER # 0 2677 # 1 2574 # Total 5251 # $percentages # D$GENDER # 0 50.98 # 1 49.02 # Total 100.00 # $validity # [1] "All tables are valid!" |
In DataSHIELD tabulated data are flagged as |
The function
ds.table2D
creates two-dimensional contingency tables of a categorical variable. Similar to
ds.table1D
the default is set to run on pooled data from all studies, to obtain an output of each study set the argument *type='split'.
DIS_DIAB
(diabetes status) and
GENDER
. The function returns the same output as
ds.table1D
but additionally computes a chi-squared test for homogeneity on (nc-1)*(nr-1) degrees of freedom (where nc is the number of columns and nr is the number of rows):ds.table2D(x='D$DIS_DIAB', y='D$GENDER') # $counts # $counts$`pooled-D$DIS_DIAB(row)|D$GENDER(col)` # 0 1 Total # 0 2625 2549 5174 # 1 52 25 77 # Total 2677 2574 5251 # $rowPercent # $rowPercent$`pooled-D$DIS_DIAB(row)|D$GENDER(col)` # 0 1 Total # 0 50.73 49.27 100 # 1 67.53 32.47 100 # Total 50.98 49.02 100 # $colPercent # $colPercent$`pooled-D$DIS_DIAB(row)|D$GENDER(col)` # 0 1 Total # 0 98.06 99.03 98.53 # 1 1.94 0.97 1.47 # Total 100.00 100.00 100.00 # $chi2Test # $chi2Test$`pooled-D$DIS_DIAB(row)|D$GENDER(col)` # Pearson's Chi-squared test with Yates' continuity correction # data: pooledContingencyTable # X-squared = 7.9078, df = 1, p-value = 0.004922 # $validity # [1] "All tables are valid!" |
It is currently possible to produce 3 types of graphs in DataSHIELD:
Scatter plots are not possible in DataSHIELD because they display individual data points and are hence potentially disclosive. Instead it is possible to draw a contour plot or a heatmap plot if the aim is to visualise a correlation pattern. |
In DataSHIELD histogram outliers are not shown as these are potentially disclosive. The text summary of the function printed to the client screen informs the user of the presence of classes (bins) with a count smaller than the minimal cell count set by data providers. |
ds.histogram
function can be used to create a histogram of
LAB_HDL
that is in the assigned variable dataframe (named
D
, by default). The default analysis is set to run on pooled data from all studies:ds.histogram(x='D$LAB_HDL') |
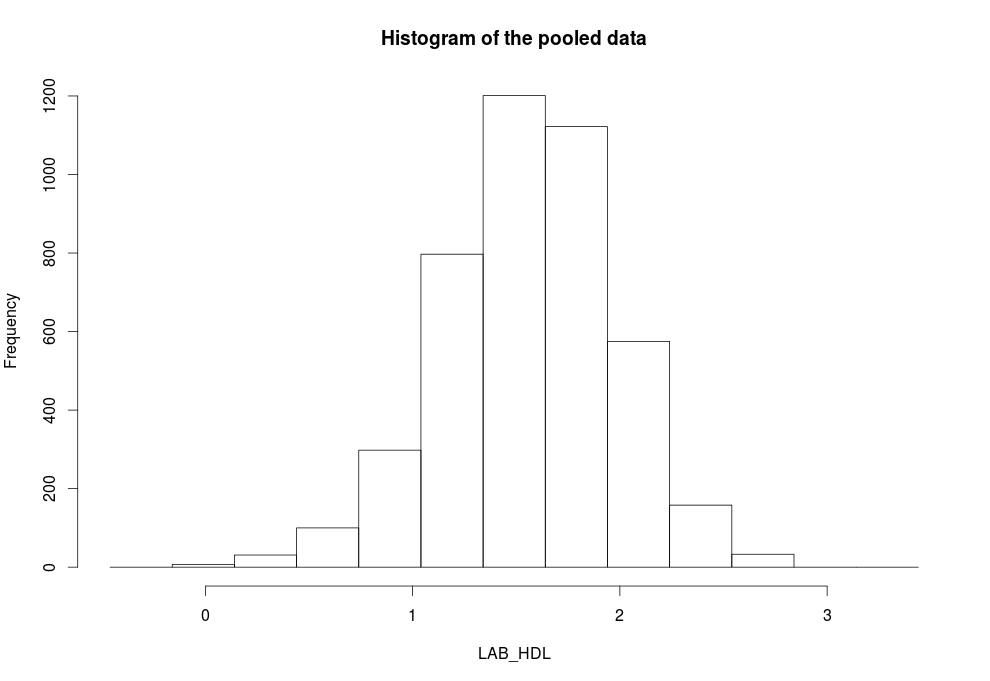
type='split'
is used. Note that some study 1 and study 2 contain 1 invalid category (bin) respectively - they contain counts less than the data provider minimal cell count:ds.histogram(x='D$LAB_HDL', type='split') #Warning: study1: 1 invalid cells #Warning: study2: 1 invalid cells |
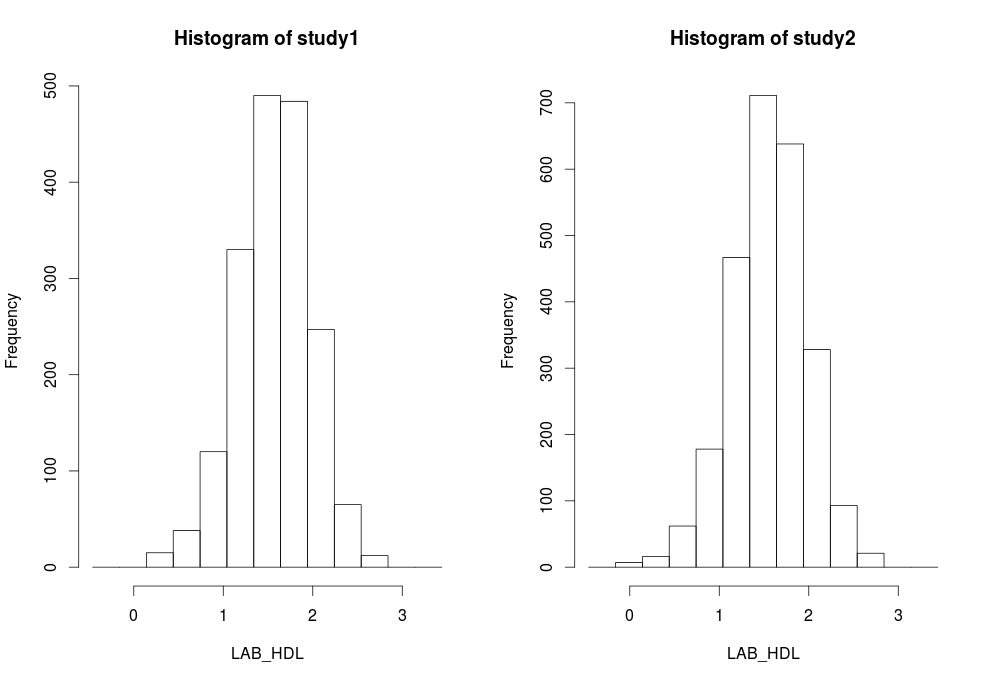
Contour plots are used to visualize a correlation pattern that would traditionally be determined using a scatter plot (which can not be used in DataSHIELD as it is potentially disclosive).
ds.contourPlot
is used to visualise the correlation between the variables
LAB_TSC
(total serum cholesterol and
LAB_HDL
(HDL cholesterol). The default is
type='combined'
- the results represent a contour plot on pooled data across all studies:ds.contourPlot(x='D$LAB_TSC', y='D$LAB_HDL') #study1: Number of invalid cells (cells with counts >0 and <5) is 83 #study2: Number of invalid cells (cells with counts >0 and <5) is 97 |
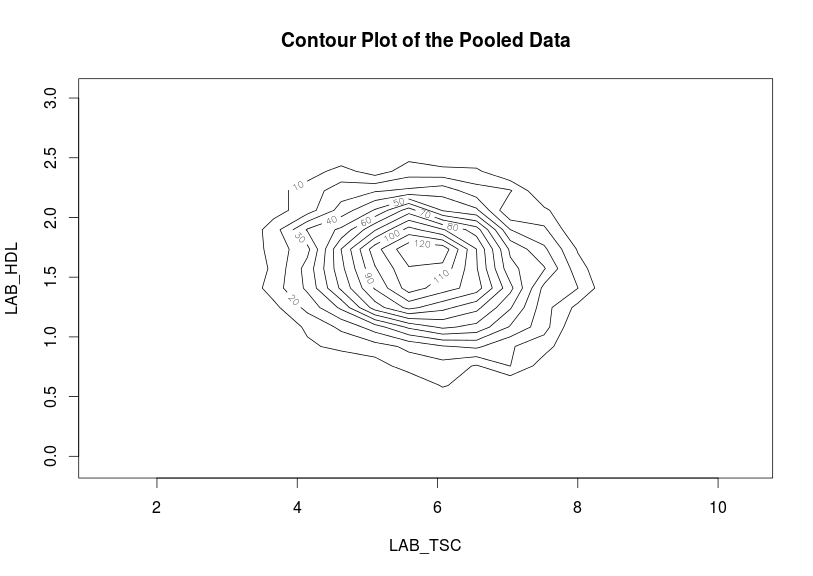
An alternative way to visualise correlation between variables is via a heat map plot.
ds.heatmapPlot
is applied to the variables
LAB_TSC
and
LAB_HDL
:ds.heatmapPlot(x='D$LAB_TSC', y='D$LAB_HDL') #study1: Number of invalid cells (cells with counts >0 and <5) is 92 #study2: Number of invalid cells (cells with counts >0 and <5) is 93 |
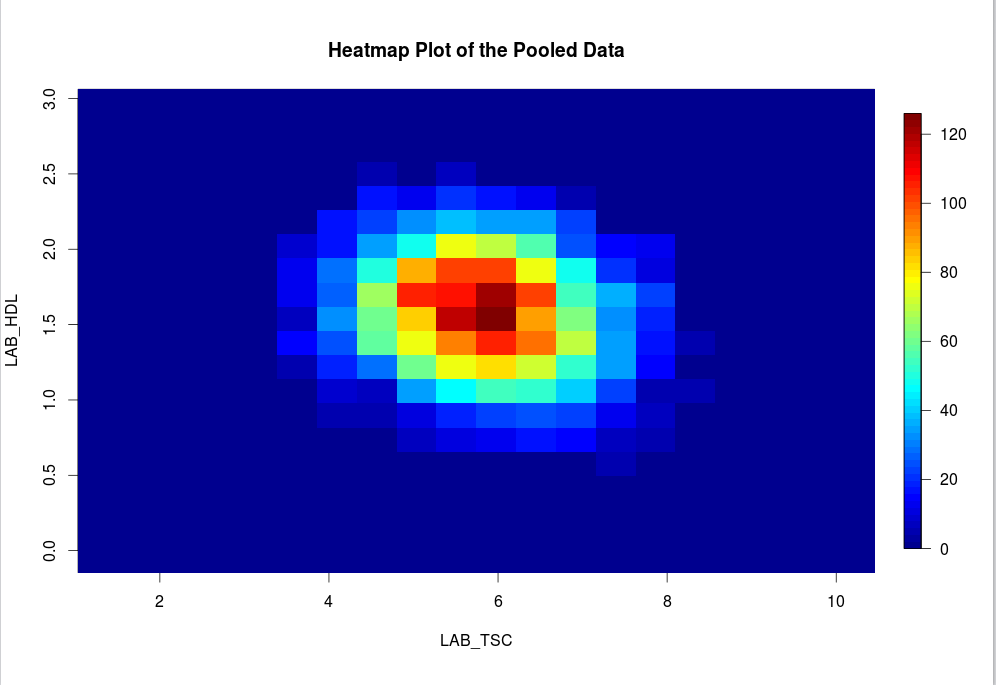
The functions ds.contourPlot and ds.heatmapPlot use the range (mimumum and maximum values) of the x and y vectors in the process of generating the graph. But in DataSHIELD the minimum and maximum values cannot be returned because they are potentially disclosive; hence what is actually returned for these plots is the 'obscured' minimum and maximum. As a consequence the number of invalid cells, in the grid density matrix used for the plot, reported after running the functions can change slightly from one run to another. In the above output for example the number of invalid cells are respectively 83 and 97 for study1 and study2. These figures can change if the command is ran again but we should not be alarmed by this as it does not affect the plot itself. It was a deliberate decision to ensure the real minimum and maximum values are not returned. |
plot tabexport > save as image 
Sub-setting is particularly useful in statistical analyses to break down variables or tables of variables into groups for analysis. Repeated sub-setting, however, can lead to thinning of the data to individual-level records that are disclosive (e.g. the statistical mean of a single value point is the value itself). Therefore, DataSHIELD does not subset an object below the minimal cell count set by the data providers (typically this is ≤ 4 observations). |
In DataSHIELD there are currently 3 functions that allow us to generate subsets data:
ds.subsetByClass
function generates subsets for each level of a
categorical
variable. If the input is a data frame it produces a subset of that data frame for each class of each categorical variable held in the data frame.
variables
argument, and the name of the subset data using the
subsets
argument.
GENDER
from our assigned data frame
D
, the subset data is named
GenderTables
:ds.subsetByClass(x = 'D', subsets = "GenderTables", variables = 'GENDER') |
ds.subsetByClass
is held in a
list
object stored server-side, as the subset data contain individual-level records. If no name is specified in the
subsets
argument, the default name of
subClasses
is used.Running ds.subsetByClass on a data frame without specifying the categorical variable in the argument |
In the previous example, the
GENDER
variable in assigned data frame
D
had females coded as 0 and males coded as 1. When
GENDER
was subset using the ds.subsetByClass function, two subset tables were generated for each study dataset
one for females and one for males.
ds.names
function obtains the names of these subset data:ds.names('GenderTables')
#$study1
#[1] "GENDER.level_0" "GENDER.level_1"
#$study2
#[1] "GENDER.level_0" "GENDER.level_1"
|
The
ds.meanByClass
function generates subset tables similar to
ds.subsetByClass
but additionally calculates the mean, standard deviation and size for each subset for specific numeric variables.
ds.meanByClass
function returns the mean, standard deviation and size of the variable defined in the
outvar
argument for each subset data defined in the
covar
argument.
LAB_HDL
(cholesterol) for the subset data
GENDER
(females and males) the default in the function is
type='combined'
(pooled across all studies). To get each study separately the argument
type='split'
must be used:ds.meanByClass(x='D', outvar='LAB_HDL', covar='GENDER') #Generating the required subset tables (this may take couple of minutes, please do not interrupt!) #--study1 # GENDER... #--study2 # GENDER... #LAB_HDL - Processing subset table 1 of 2... #LAB_HDL - Processing subset table 2 of 2... # D.GENDER0 D.GENDER1 #LAB_HDL(length) "2677" "2574" #LAB_HDL(mean&sd) "1.51(0.44)" "1.62(0.4)" |
The |
The function
ds.subset
allows general sub-setting of different data types e.g. categorical, numeric, character, data frame, matrix. It is also possible to subset rows (the individual records). No output is returned to the client screen, the generated subsets are stored in the server-side R session.
ds.subset
to subset the assigned data frame
D
by rows (individual records) that have no missing values (missing values are denoted with
NA
) given by the argument
completeCases=TRUE
. The output subset is named
D_without_NA
:ds.subset(x='D', subset='D_wihthout_NA', completeCases=TRUE) #In order to indicate that a generated subset dataframe or vector is invalid all values within it are set to NA! |
The |
D
with BMI values ≥ 25 using the argument
logicalOperator
. The subset object is named
BMI25plus
using the
subset
argument and is not printed to client screen but is stored in the server-side R session:ds.subset(x='D', subset='BMI25plus', logicalOperator='PM_BMI_CONTINUOUS>=', threshold=25) #In order to indicate that a generated subset dataframe or vector is invalid all values within it are set to NA! |
The subset of data retains the same variables names i.e. column names |
ds.quantileMean
with the argument
type='split'
will confirm the BMI results for each study are ≥ 25.ds.quantileMean('BMI25plus$PM_BMI_CONTINUOUS', type='split')
# $study1
# 5% 10% 25% 50% 75% 90% 95% Mean
# 25.3500 25.7100 27.1500 29.2000 32.0600 34.6560 36.4980 29.9019
# $study2
# 5% 10% 25% 50% 75% 90% 95% Mean
# 25.46900 25.91800 27.19000 29.27000 32.20500 34.76200 36.24300 29.92606
|
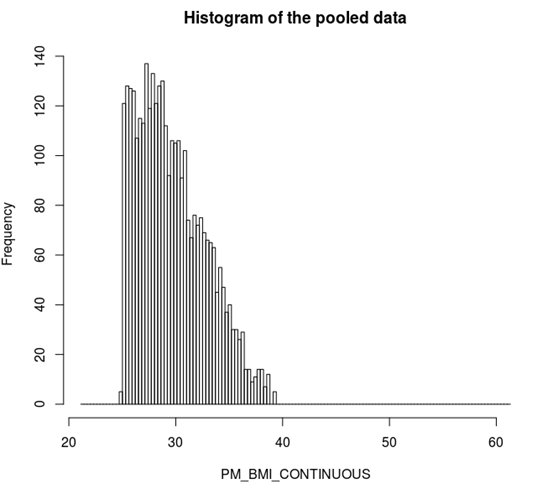
ds.histogram('BMI25plus$PM_BMI_CONTINUOUS') |
Horizontal DataSHIELD allows the fitting of:
In GLM function the outcome can be modelled as continuous, or categorical (binomial or discrete). The error to use in the model can follow a range of distribution including gaussian, binomial, Gamma and poisson. In this section only one example will be shown, for more examples please see the manual help page for the function.
The function
ds.glm
is used to analyse the outcome variable
DIS_DIAB
(diabetes status) and the covariates
PM_BMI_CONTINUOUS
(continuous BMI),
LAB_HDL
(HDL cholesterol) and
GENDER
(gender), with an interaction between the latter two. In R this model is represented as
D$DIS_DIAB ~ D$PM_BMI_CONTINUOUS+D$LAB_HDL*D$GENDER |
By default intermediate results are not printed to the client screen. It is possible to display the intermediate results, to show the coefficients after each iteration, by using the argument
display=TRUE
.
ds.glm(formula=D$DIS_DIAB~D$PM_BMI_CONTINUOUS+D$LAB_HDL*D$GENDER, family='binomial') #Iteration 1... #CURRENT DEVIANCE: 5772.52971970323 #Iteration 2... #CURRENT DEVIANCE: 1316.4695853419 .... #SUMMARY OF MODEL STATE after iteration 9 #Current deviance 534.369101004808 on 4159 degrees of freedom #Convergence criterion TRUE (1.38346480863211e-12) #beta: -6.90641416691308 0.142256253558181 -0.96744073975236 -1.40945273343361 0.646007073975006 #Information matrix overall: # (Intercept) PM_BMI_CONTINUOUS LAB_HDL GENDER1 #(Intercept) 52.47803 1624.8945 71.78440 16.77192 #PM_BMI_CONTINUOUS 1624.89450 51515.6450 2204.90085 503.20484 #LAB_HDL 71.78440 2204.9008 109.48855 25.91140 #GENDER1 16.77192 503.2048 25.91140 16.77192 #LAB_HDL:GENDER1 25.91140 774.1803 42.87626 25.91140 # LAB_HDL:GENDER1 #(Intercept) 25.91140 #PM_BMI_CONTINUOUS 774.18028 #LAB_HDL 42.87626 #GENDER1 25.91140 #LAB_HDL:GENDER1 42.87626 #Score vector overall: # [,1] #(Intercept) -3.610710e-10 #PM_BMI_CONTINUOUS -9.867819e-09 #LAB_HDL -6.308052e-10 #GENDER1 -2.910556e-10 #LAB_HDL:GENDER1 -5.339693e-10 #Current deviance: 534.369101004808 #$formula #[1] "D$DIS_DIAB ~ D$PM_BMI_CONTINUOUS + D$LAB_HDL * D$GENDER" #$family #Family: binomial #Link function: logit # #$coefficients # Estimate Std. Error z-value p-value low0.95CI.LP #(Intercept) -6.9064142 1.08980103 -6.3373166 2.338013e-10 -9.04238494 #PM_BMI_CONTINUOUS 0.1422563 0.02932171 4.8515676 1.224894e-06 0.08478676 #LAB_HDL -0.9674407 0.36306348 -2.6646601 7.706618e-03 -1.67903208 #GENDER1 -1.4094527 1.06921103 -1.3182175 1.874308e-01 -3.50506784 #LAB_HDL:GENDER1 0.6460071 0.69410419 0.9307062 3.520056e-01 -0.71441214 # high0.95CI.LP P_OR low0.95CI.P_OR high0.95CI.P_OR #(Intercept) -4.7704434 0.00100034 0.0001182744 0.008405372 #PM_BMI_CONTINUOUS 0.1997257 1.15287204 1.0884849336 1.221067831 #LAB_HDL -0.2558494 0.38005445 0.1865544594 0.774258560 #GENDER1 0.6861624 0.24427693 0.0300447352 1.986079051 #LAB_HDL:GENDER1 2.0064263 1.90790747 0.4894797709 7.436693232 #$dev #[1] 534.3691 #$df #[1] 4159 #$nsubs #[1] 4164 #$iter #[1] 9 #attr(,"class") #[1] "glmds" |
After every iteration in the glm, each study returns non disclosive summaries (a score vector and an information matrix) that are combined on the client R session. The model is fitted again with the updated beta coefficients, this iterative process continues until convergence or the maximum number of iterations is reached. The output of |
You have now sat our basic DataSHIELD training. Remember you can:
|